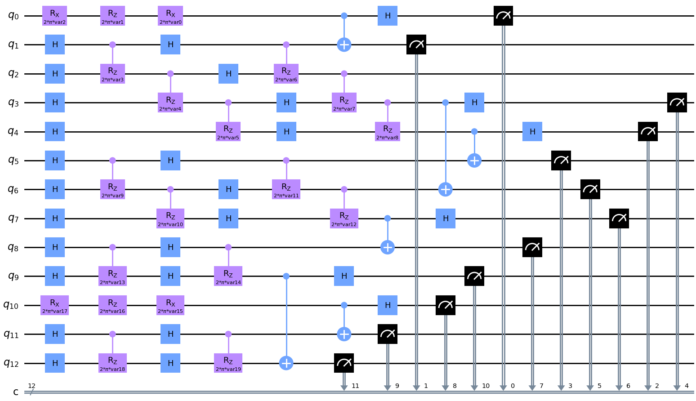
Cambridge Quantum has unveiled what is touted as the world’s first toolkit and library for Quantum Natural Language Processing (QNLP), named lambeq after the late mathematician and linguist Joachim Lambek.
Capable of converting sentences into a quantum circuit, lambeq is designed to accelerate the development of practical, real-world QNLP applications, such as automated dialogue, text mining, language translation, text-to-speech, language generation and bioinformatics.
lambeq has been released on a fully open-sourced basis for the benefit of the world’s quantum computing community and the rapidly growing ecosystem of quantum computing researchers, developers and users.
lambeq works seamlessly with CQ’s TKET quantum software development platform that is also fully open-sourced. This provides QNLP developers with access to the broadest possible range of quantum computers.
CQ’s Oxford-based quantum computing research team that conceived, designed and engineered lambeq was led by Chief Scientist Bob Coecke, with senior scientist Dimitrios Kartsaklis, as chief architect of the platform.
“The release of lambeq is the natural next step after the publication a few months ago that provided details of the world’s first QNLP implementation by CQ on actual quantum computers, and our initial disclosure of the foundational principles in December 2019,” said Coecke.
lambeq removes the barriers to entry for practitioners and researchers who are focused on AI and human-machine interactions, potentially one of the most significant applications of quantum technologies. TKET has gained a worldwide user base now in the hundreds of thousands.
The toolkit has the potential to become the most important toolkit for the quantum computing community seeking to engage with QNLP applications that are amongst the most important markets for AI.
A key point that has become apparent recently is that QNLP will also be applicable to the analysis of symbol sequences that arise in genomics as well as in proteomics.